Aging is a highly complex process with thousands of genes influencing our health, which poses a challenge for researchers looking to explain and target the underlying processes that lead to declining health. Researchers from the Babraham Institute’s Epigenetics research program have published a map of genetic interactions in C. elegans in iScience which can be used to identify new genes that influence lifespan and that have equivalent genes in humans.
Researchers use simple model organisms like the nematode worm C. elegans to gather information that can inform studies on human aging because many genes are shared or have counterparts in other species. However, there are some conceptual and technical challenges that apply to the study of aging in model organisms. Dr. Casanueva, Group leader in the Epigenetics research program explains: “The way researchers usually study gene function is by disrupting its function and observing what happens. The disruption of some genes causes worms to live a very long-life. In this way, researchers have found the so-called ‘longevity-pathways.” However, the complexity underlying aging means that it is not enough to focus on individual genes. We need to study the overall organization of longevity by generating a systems-wide view.”
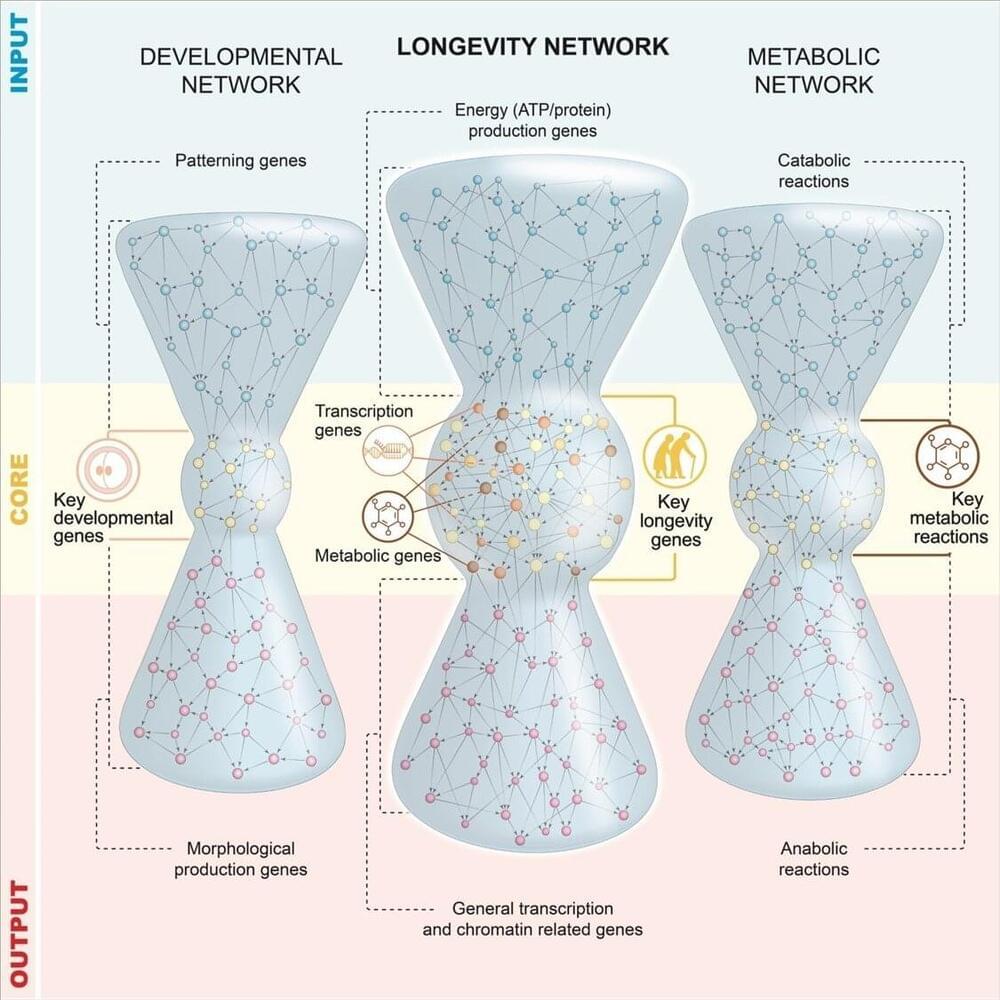
In collaboration with the physicist Marta Sales Pardo at University of Rovira i Virgili, Dr. Casanueva and her lab set out to cast a wider net when it comes to studying longevity genes. Together they created the largest network of gene regulatory interactions that are found in a long-lived type of C. elegans. In this network, the relationships between genes are represented by lines, and represented in different layers based on the flow of information between genes. The middle of the web represents the genes with the most influence, in this case, they receive complex input signals and de-code them, and connect to an output layer of genes. The researchers found that most key genes for longevity belong to transcription factors and metabolic genes.